I built a blood work lab result analyzer app, and the process was really fun. Want to share my thoughts and approach, hoping this can be inspiring and help you, my friend, to utilize AI in our day-to-day life. Below is the screen recording of me using the app to analyze some lab results.
Opening a lab report feels like trying to read ancient hieroglyphs. You see things like “Hypersegmented Neutrophils” or “Mean Corpuscular Volume,” and suddenly you’re spiraling on WebMD, Google searching, YouTube Dr.Berg, jumping between the medical portal to copy the biomarker and paste online to see what it means, how to improve it, too much work and anxiety
I decided to make a fun project just for myself to skip all this tedious research and have a centralized place to provide explanations in layman’s language, and a wellness plan.
Using Python, Claude AI, Prompt Engineering, and a dash of coding magic, it turns those scary PDFs into simple, “explain-it-to-me-like-I’m-five” health insights. Here’s how I did it (and how you can, too!)
The Vision: A Health Assistant in Your Pocket
The goal was simple:
- Upload a messy PDF.
- Translate the science-speak into plain English.
- Generate a “What’s Next?” plan so you actually know what to eat or do differently.
My “Secret Weapon”: Claude Code
Instead of staring at a blank screen for hours, I used Claude Code—a terminal-based AI assistant. It felt like having a senior developer sitting right next to me. I just told the terminal what I wanted, and bam!—it scaffolded the project, set up my virtual environment, and wrote the boilerplate code.
If you’re on a Mac, the setup is a total breeze. A few brew install commands and you’re living in the future.
The Recipe (The Tech Stack)
- Streamlit: For the “Face” of the app. It makes Python look like a professional website in seconds.
- pdfplumber: My “eyes.” It digs through those stubborn PDF tables to find the numbers that matter.
- Claude 3.5 Sonnet: The “Brain.” It takes the raw data and provides the supportive, layman-friendly explanations.
- Python-dotenv: The “Vault.” It keeps my API keys safe and secret.
Step-by-Step: The Build
- The Scaffold: I let Claude Code generate the directory and the basic
app.py. - The Extraction: We taught the app to identify “Reference Ranges.” If your Glucose is 115 but the range stops at 99, the app flags it!
- The Prompt: This was the fun part. I told Claude to act like a “Friendly Health Guide.” No alarmist tones allowed—just helpful facts and clear action steps.
- The Test: I generated a dummy report (shoutout to
fpdf!) with high cholesterol and low Vitamin D just to see if the AI would catch it. It did! It suggested more sunshine and some heart-healthy fats.
Detailed steps:
- Create your project directory in your terminal
mkdir blood-work-analyzer && cd blood-work-analyzer
2. Launch Claude Code from your terminal
Claude
3. Prompt – This does take some engineering knowledge
below was what I input:
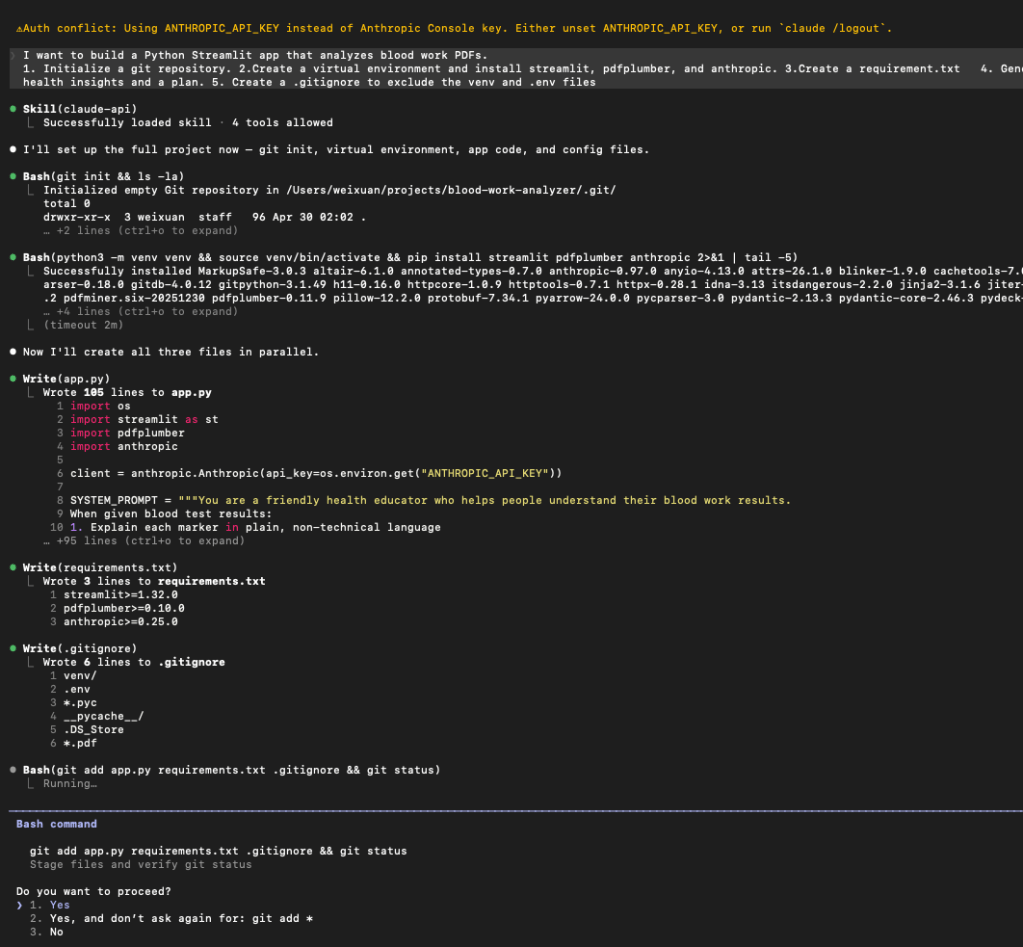
I want to build a Python Streamlit app that analyzes blood work PDFs. Create a .gitignore to exclude the venv and .env files. Initialize a git repository. Create a virtual environment and install streamlit, pdfplumber, and anthropic. Create a requirements.txt. Generate a app.py that handles PDF uploads, extracts text using pdfplumber, and uses the Claude API to provide layman’s health insights and a plan.
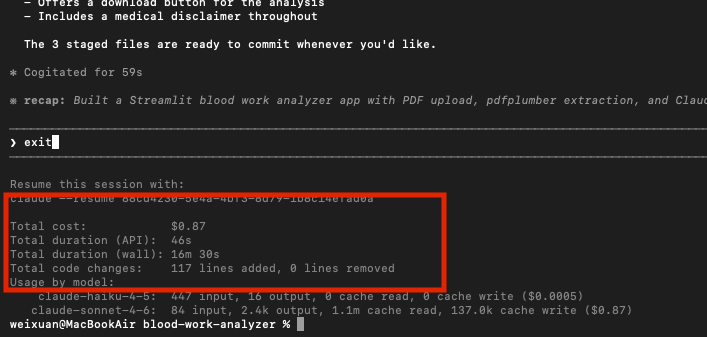
I was sweating when I saw it start using more and more tokens, I was worried about the API billing, but it was not bad at all. the entire project only used 21k tokens and cost me 0.87$
fantastic now to make it have a certain tune, in the application app.py code, need to slightly modify the role/tune prompt. you do what you feel like. Below was mine.
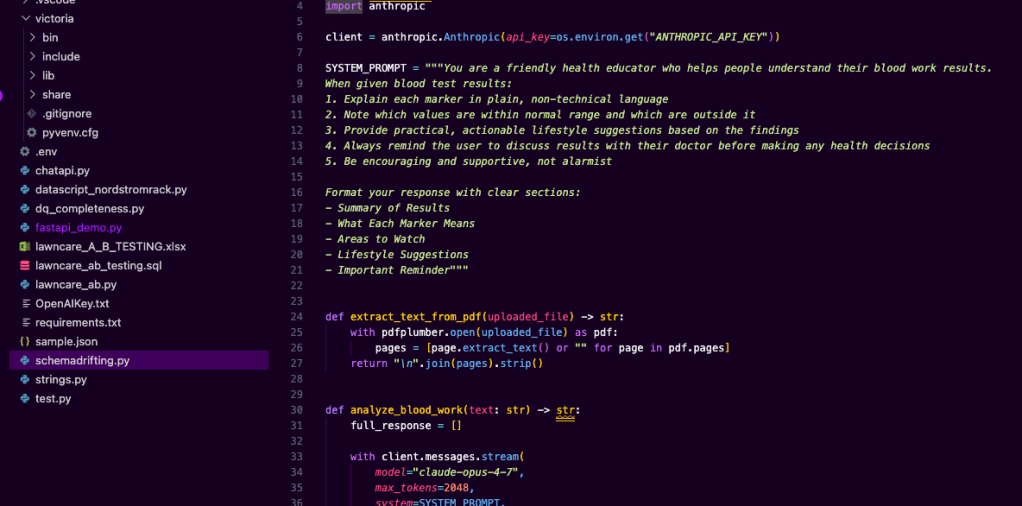
4. Set up API key so your app can run the ai model. create a .env file in your project directory, add the API key created from Claude Console. put
ANTHROPIC_API_KEY="your_key_here"
And now you can activate your environment, install the libraries, and launch the app. First-time launch of Streamlit in the terminal, it will ask for your email. You don’t have to provide it; just press enter and continue. The terminal will output a URL (usually http://localhost:8501). Your default web browser should open this automatically
source venv/bin/activatepip install streamlit pdfplumber anthropic python-dotenvstreamlit run app.py
5. Testing the workflow
Once the browser window opens:
- Upload: Drag and drop a PDF of a lab report.
- Processing: Watch the terminal; if there are errors in the Python code, they will appear there in red text.
- Review: Ensure the “Layman Language” section is easy to read and the “Action Plan” is formatted correctly.
Tip: If the app crashes or the PDF doesn’t parse correctly, you can go back to your Claude Code terminal and say: “The app is giving an error on line 42 when I upload a PDF. Can you fix the pdfplumber logic to handle multi-page tables?
Takeaways
Building this wasn’t just about the code; it was about empowerment. Technology is at its best when it takes something complex and makes it accessible. Now, instead of being intimidated by a lab report, I have a tool that helps me prepare for my next doctor’s visit with confidence.
Want to see the code? Check out my project on GitHub! (Link below)

Leave a comment